Standalone neuron_s example with explicit connectivity.
- Author
- Oscar Sotomayor
This file provides a minimal, executable example that demonstrates how multiple neuron_s instances can be connected and evaluated without using the full NeuroTIC network container.
The example:
- constructs a small set of neurons manually,
- connects them using direct pointer references,
- evaluates them using an explicit external loop,
- and prints the resulting output.
The purpose of this file is illustrative. It shows how neuron_s behaves as a passive structure when driven by user-defined execution code.
This example does not represent the complete NeuroTIC execution model. No net_s, wiring_s, training, or activation dispatch is involved.
For the structural definition of neuron_s, see:
Example topology
The following diagram represents the concrete topology implemented in this example.
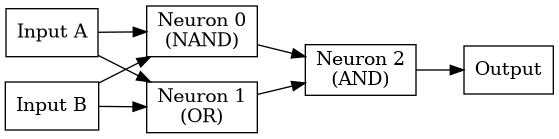
/
#define NEURON_0 NAND
#define NEURON_1 OR
#define NEURON_2 AND
#define AND .w= (weight_t []){ 0.1 , 0.1 } , .b= (bias_t)(-0.15)
#define NAND .w= (weight_t []){ -0.1 , -0.1 } , .b= (bias_t)( 0.15)
#define OR .w= (weight_t []){ 0.1 , 0.1 } , .b= (bias_t)(-0.05)
#define NOR .w= (weight_t []){ -0.1 , -0.1 } , .b= (bias_t)( 0.05)
#define CONJUNCION_A .w= (weight_t []){ -0.1 , 0.1 } , .b= (bias_t)(-0.05)
#define CONJUNCION_B .w= (weight_t []){ 0.1 , -0.1 } , .b= (bias_t)(-0.05)
#define IMPLICATION_A .w= (weight_t []){ -0.1 , 0.1 } , .b= (bias_t)( 0.05)
#define IMPLICATION_B .w= (weight_t []){ 0.1 , -0.1 } , .b= (bias_t)( 0.05)
#include <stdio.h>
int main( void ) {
data_t inputs[2];
[0]= { .
inputs=
sizeof( inputs ) /
sizeof( data_t ) , .bff_idx= 0 , .in= (data_t *[]){ &inputs[0] , &inputs[1] } , NEURON_0 , .fn= 0 , .out= (data_t)(0.0) },
[1]= { .inputs=
sizeof( inputs ) /
sizeof( data_t ) , .bff_idx= 0 , .in= neuron[0].
in , NEURON_1 , .fn= 0 , .out= (data_t)(0.0) },
[2]= { .inputs= 2 , .bff_idx= 0 , .in= (data_t *[]){ &neuron[0].
out , &neuron[1].
out } , NEURON_2 , .fn= 0 , .out= (data_t)(0.0) }
};
for( uint8_t i= 0 ; i < 4 ; i++ ){
for( size_t j= 0 , f= sizeof( inputs ) / sizeof( data_t ) ; j < f ; j++ ) inputs[j]= !!(data_t)( i & ( 1 << j ) );
for(
size_t j= 0 , f=
sizeof( neuron ) /
sizeof(
neuron_s ) ; j < f ; j++ ){
for( input_t k= 0 ; k < neuron[j].
inputs ; k++ ) sum+= *neuron[j].in[k] * neuron[j].w[k];
neuron[j].
out= (data_t)( sum >= 0.0f );
}
printf( "| input_A: %0.0f | input_B: %0.0f | output: %0.0f |\n", inputs[0] , inputs[1] , neuron[2].out );
}
return 0;
}
Fundamental structural definitions of the NeuroTIC runtime model.
Structural unit representing a computation node.
Definition ntcore.h:56
data_t ** in
Definition ntcore.h:59
input_t inputs
Definition ntcore.h:57
data_t out
Definition ntcore.h:63
bias_t b
Definition ntcore.h:61
Definition in file neurons.c.
